The one-year funded period for SageCite came to an end in July 2011. It took a further few weeks to tie up some loose ends, and follow up on some of our dissemination activities. As a belated sign off on the project, this post sums up the outputs of SageCite.
The Deliverables section of this blog provides links to all the concrete outputs of the project, hosted both on this blog and outside it. The latter include the KnowledgeBlog instance that hosts reviews of technologies and opinion pieces on citation, and the Benefits Analysis of citation in the domain of disease network modelling, using the KRDS methodology. The deliverables page also brings together the write-up on the Taverna demonstrator developed by SageCite and the description of the application domain. Finally, the publisher interviews are also linked there.
The Dissemination page has been updated with all the talks in which SageCite has featured as well as presentations that continued beyond the funded period. We have also listed other events attended by the project team, as these will be of wider interest to the community.
Less tangible as an outcome has been the interaction with the data citation community, which has grown over the lifetime of the project. The community spans both standard or service -focussed activities, such as ORCID and DataCite, and more generic meetings like Beyond The PDF and the International Symposium. It has been a privilege for the members of the SageCite team learn from and share insight with this inspiring community. A summary of all the activity around data citation was written up in an editorial for the Digital Preservation Coalition newsletter in December 2011.
The last in my trio of belated blog posts is based on the report: Riding the wave – How Europe can gain from the rising tide of scientific data – Final report of the High Level Expert Group on Scientific Data, published in October 2010.
Once again I have extracted from this report some content that seems most relevant to discussions of data citation and attribution. First up is noting that “Develop and use new ways to measure data value, and reward those who contribute it” (P. 5-6. Point 3) was selected as one of six in a short-list of actions for EU institutions, listed in the Executive summary:
“If we are to encourage broader use, and re-use, of scientific data we need more, better ways to measure its impact and quality. We urge theEuropean Commission to lead the study of how to create meaningful metrics, in collaboration with the ‘power users’ in industry and academia, and in cooperation with international bodies.”
In scenarios of the future, scenario 3 (p. 14) describes how an academic creates a cleaned results data set and makes it publicly available; she imagines the result set becoming as popular as Top-40 song, with the consequence that “her chances for tenure rise”. The report goes on to explore incentives for contributing data:
“How can we get researchers – or individuals – to contribute to the global data set? Only if the data infrastructure becomes representative of the work of all researchers will it be useful; and for that, a great many scientists and citizens will have to decide it is worth their while to share their data, within the constraints they set. To start with, this will require that they trust the system to preserve, protect and manage access to their data; an incentive can be the hope of gain from others’ data, without fear of losing their own data. But for more valuable information, more direct incentives will be needed – from career advancement, to reputation to cash. Devising the right incentives will force changes in how our universities are governed and companies organised. This is social engineering, not to be undertaken haphazardly.” (P. 19)
The report then describes some milestones that need to be achieved to realise their vision of a data infrastructure for 2030, where data is a valuable asset and the infrastructure supports ‘seamless access, use, re-use, and trust of data’, and the expected impact of each milestone. One of the main milestones in this vision is that of producers of data who share it openly, and with confidence in the sharing infrastructure. The expected impact of this milestone is predicted to be “Researchers are rewarded, by enhanced professional reputation at the very least, for making their data available to others. Confidence that their data cannot be corrupted or lost reassures them to share even more. Data sharing, with appropriate access control, is the rule, not the exception. Data are peer-reviewed by the community of researchers re-using and re-validating them. The outcome: A data-rich society with information that can be used for new and unexpected purposes.” The report also describes the risk of inaction if that milestone is not achieved: “Information stays hidden. The researcher who created it in the hope it can yield more publications or patents in the future holds on to it. Other researchers who need that information are unable to get at it, or waste time re- creating it. The outcome: A world of fragmented data sources – in fact, a world much like today.” (P.25)
As mentioned earlier, in their call to action, the report writers selected Develop and use new ways to measure data value, and reward those who contribute it as one of six first steps that need to be acted upon. The need for universal metrics is described – although the report stops short of exploring the potential role for data linking and citation in the developing of these metrics.
“Who contributes the most or best to the data commons? Who uses the most? What is the most valuable kind of data – and to whom? How efficiently is the data infrastructure being used and maintained? These are all measurement questions. At present, we have lots of different ways of answering them – but we need better, more universal metrics. If we had them, funding agencies would know what they are getting for their money – who is using it wisely. Researchers would know the most efficient pathways to get whatever information they are seeking. Companies would be able to charge more easily for their services.” (P. 32)
The second of the three posts that I’m releasing on the principle of ‘better late than never’ was a pointer to Christine Borgman’s report: Research Data: Who Will Share What, with Whom, When, and Why?“ (2010), which was summarised by Current Cites as follows:
Summary by Current Cites : The growing open data and open science movements have helped focus attention on issues related to scientific research data. In this eprint, Borgman, Presidential Chair and Professor of Information Studies at UCLA, tackles the thorny problem of defining “data,” examines the purposes of data-driven research, discusses the methods of such research, summarizes researchers’ incentives and disincentives for data sharing, takes a detailed look at four policy arguments for data sharing, and considers the role of libraries in the data sharing process. The data-sharing policy arguments are “to make the results of publicly funded data available to the public, to enable others to ask new questions of extant data, to advance the state of science, and to reproduce research. http://lists.webjunction.org/currentcites/2010/cc10.21.9.html
Borgman, who chaired the International Symposium and Workshop on Developing Data Attribution and Citation Principles and Practice, gave an excellent summary of the state of data citation in her opening talk. In the above report, she discusses what could motivate or induce researchers to share data “Incentives for researchers to share their data include the ethos of open science and peer review; the value of collaborating with others, for which data may be the “glue;” benefits to reputation; and reciprocity. Depositing one’s data may be a condition of gaining access to the data of others, and of access to useful tools for analysis and management. Coercion may also play a role: some funding agencies or individual grant contracts may require data contribution as a condition for funding.” (P.7) and the key citation-related requirement that ”At a minimum, most researchers want attribution for any data used by others.” (P.9)
Borgman also comments that “Learning the interests of a given community, however narrowly or broadly defined, requires close engagement and study.” (P.14) – this was very pertinent to the work of SageCite, which focussed on a specific community at Sage Bionetworks, and put effort into learning about and reproducing their workflows as a step towards understanding the data citation needs.
In the process of tying up loose ends for the SageCite project, I’ve unearthed three draft blog postings, all of which were related to other blog posts or publications which comment on data sharing, re-use or citation – three key themes for the SageCite project. All of them are still very relevant despite the time that has passed since I first came across them, so I’ve decided to let them see the light of day.
The first was a blog post over on Jeni Tennison’s blog. Jeni was very active in efforts to release UK government data as open linked data and in this post from September 2010 she reflected on what it would take to ensure the public sector would ‘publish reusable data in the long term’. It struck me that this quote from Jeni’s post about releasing government data as open data could apply to the publishing of scientific research data (replace ‘public sector’ with ‘research sector’ or ‘academic’). This is what Jeni had to say about the ease of releasing government data, and reward systems:
“To do that, data publication needs to be sustainable. It needs to be embedded within the day-to-day activity of the public sector, something that seems as natural as the generation of PDF reports seems today. It also needs to be useful. It needs to be easy for anyone to understand and reuse the data, with minimal effort. It cannot be the case, long term, that you need to be an expert hacker to reuse government data. …….
To get there, we need to work towards a virtuous cycle in which the public sector is rewarded for publishing useful data well. The reward may come from financial savings, from increasing data quality, from better delivery of its remit, or simply from kudos. It doesn’t matter how, but there needs to be some reward, or it just won’t happen.“
I’ve just described the background to this post in the previous blog entry. So without further ado:
Requirement 1. The Citation needs to be able to uniquely identify the object cited.
This seemingly simple requirement is about making sure that the citation contains an unambigous reference to the cited object. However unpacking this requirement reveals some difficult consequences. Textual citations can be made unique through a combination of fields e.g. author, title, date. Truly unambiguous identification could alternatively be achieved through the use of unique identifiers. Do all the different types of citable object need to be identified and listed? How does a discipline work out and agree the granular level at which objects can be (or need to be) identified (ie cited), whether uniquely or as a package.
Requirement 2. The Citation needs to support the retrieval of the cited object.
Location of cited objects is an important driver for citation since it supports re-use, validation and reproducibility. Retrieval can be achieved indirectly by providing sufficient information for a human user to process the parts of the citation and use knowledge of the discipline (e.g. location of discipline-specific repositories and methods to search and access their contents) to find the cited object. This requirement also subsumes some other requirements commonly associated with identifiers e.g. persistence and implies properties of other infrastructure for data management (e.g. curation of data). Supporting automated retrieval leads us to requirement 3.
Requirement 3. The citation mechanism must be compatible with Web infrastructure.
With increasing calls for publications to present data in a form that is more interactive, any modern infrastructure for citation should be compatible with web infrastructure. In practice this implies that citations must contain actionable URLs. Any internal identifiers used for internal management must eventually be mapped to an actionable citation, which throws up interesting questions on the management of identifiers and objects as they move across curation boundaries from the private sphere to the public domain.
Requirement 4. The citation ‘system’ must be able to generate a citation with all the desired fields.
The desired fields must be agreed for all the types of citable object. Does this really need to happen up front? How necessary is it to agree common fields and labels within and across disciplines? A number of dependencies can be identified: The information required for the citation must either be captured in the datasets (from where it can be extracted), or in the metadata, or explicitly entered by the user. Systems to automatically generate or capture metadata (for example, provenance metadata) to decrease load on user are needed. This information must be available at the point at which the citation is generated and shared – this may be internally, where a data contributor cites their own work, or within external systems when others cite a dataset.
Requirement 5 The citation mechanism must be identifier-agnostic.
Globally, the community would need to identify which identifiers are in use, what are their life-cycles? How are they used? The range could include discipline-specific identifiers and globally agreed schemes. The requirement throws up some dependencies with respect to accommodating different resolution mechanisms.
Requirement 6 The citation mechanism must support gathering of metrics
Credit and attribution are considered important drivers for citation. The computation of metrics to measure value of contrbutions is made possible through citation. Does this requirement imply that the citation must be processable, preferably in automated format? What are the implications for the other requirements, especially 1, 3 and 4.
Requirement 7 The citation must be human readable
Humans use human judgement to decide whether the cited object is worthy of more attention, if it can be trusted, and other similar judgements, which are based in part on the information within the citation. For example the mention of a trusted disciplinary repository lends confidence that the item will be retrievable and formatted to a high standard. Information about the data (e.g. species) suggests if it is within the field of interest of the reader. The name of a data contributor can also be used as a proxy for determining if data is of interest. Citations for data may lead the human reader indirectly to supplementary information (metadata) that helps to arrive at these judgements.
Requirement 8 The citation must be machine processable
This requirement is linked to several of the others. In a Linked Data environment, citations that are compatible with Web Infrastructure are required to re-use conventions and ontologies established by Linked Data practices. The automation of metric computation could be made more reliable through agreed ways of making citations machine processable. Automated following of links and gathering of information to answer questions such as ”Find me all the data contributed by contrubutor x” are requirements that future citation infrastructure should be able to meet.
I have been working on a How To guide on Data Citation which is co-authored with Alex Ball and will be published shortly by the DCC. The How-To guide is a sister publication to the Briefing Paper which was released recently. In the process of working together with Alex I provided him with a few pointers. One of the pointers was to a paper I had prepared for the 2010 Sage Congress.
I am digging out that paper and sharing it now for two reasons. Firstly, in one of the planning telcons for the Data Attribution and Citation Practices and Standards Symposium taking place this week, someone asked if we knew of a resource that describes the landscape of data citation – an introduction to various standards, technologies and initiatives. I originally went looking for this paper for a different reason (which I will be coming to in a minute) but having cast a quick eye over it, I realised I should offer it to the meeting I’m about to attend since it covers some of the ground that was requested. I have already provided links to the SageCite KnowledgeBlog which was meant to describe the data citation landscape in a more methodical way. The KnowledgeBlog would require take-up by the community to make it more comprehensive, as its current scope is too limited. The 2010 paper is slightly out of date. Neither is perfect, but together they are reasonably complementary as it happens!
But I digress. At the Harvard Data Citation Principles meeting I had suggested that as a community we should be able to agree some high level principles for data citation, however the difficulty would be in checking out the detail for each intended application area. This week’s Developing Data Attribution and Citation Practices and Standards symposium seems to be addressing just that question, looking at the practice of citation across different disciplines and asking where common ground and effort can pay off. At the Harvard meeting I was asked if I had a set of high-level princiles that we should all be able to agree on. At the time I forgot about the ones I had written into the paper for the 2010 Sage Congress, and I might not have remembered that they exist if Alex hadn’t spotted them and included them in the draft of the DCC How To guide. I guess I am still a software developer at heart as I framed that list as requirements rather than principles, and I had an eye to helping Sage Bionetworks consider requirements for the platform it was developing.
So the next post will list those requirements, mainly so they can be shared for discussion within the Data Citation community.
The SageCite project has been applying the KRDS Benefits Analysis Toolkit as a method of evaluating the benefits of data citation. We have been collaborating with the JISC-funded KRDS/I2S2 Digital Preservation Benefit Analysis Tools Project providing a case study of the application of the Benefits Framework and helping to feed into the evolution of the tool and its documentation.
Last week we presented at the successful end of project dissemination workshop which provoked lively discussion on implementing the toolkit with funders and other attendees. A report on that workshop has now been published, with a link to the presentation by the SageCite case study.
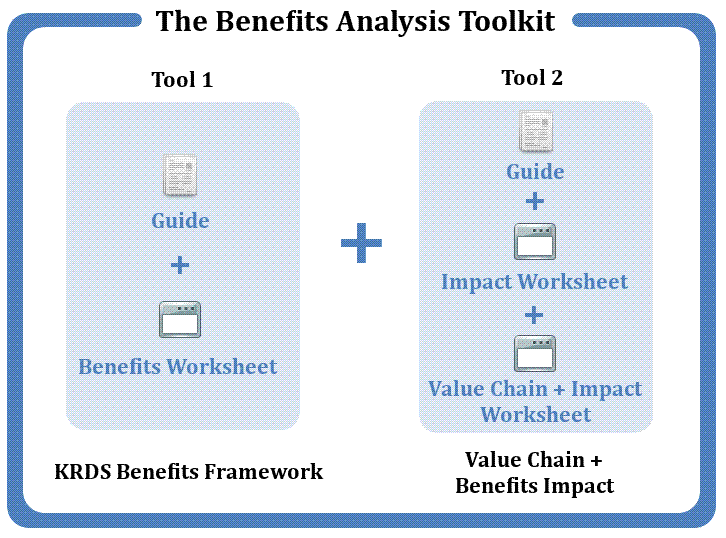
The toolkit consists of two tools: the KRDS Benefits Framework and the Value-chain and Benefits Impact Tool.
The KRDS Benefits Framework which has been tested by SageCite is an “entry-level” tool for identifying, assessing and communicating the benefits from investing resources in the curation of research data. It helps to articulate the benefits and can be customised. The benefits are organised around three dimensions: “what”, “when” and “who” of the value proposition of the activities.
Images are Copyright Charles Beagrie Limited 2011 and KRDS/I2S2 Benefits Project Partners 2011.
We will be publishing the benefits analysis carried out by SageCite after some further validation. The updated Benefits Analysis Toolkit has an official release date of 31st July and will be releases on the KRDS/I2S2 project site.
The aims of the SageCite project included extending Taverna to incorporate a citation service, and collecting evidence of the types of network models, data and process used in the modelling of disease networks. Review of the literature shows that current conventions for reporting research, and methods of citing and linking data in publications, do not always provide sufficient detail and information to reproduce the research reported. This has led to calls to change methods of publication to improve the links and citations to the research underlying the journal article. Eric Schadt described the launch of a new journal Open Network Biology to address this problem, and Phil Bourne, winner of the 2010 Jim Gray eScience award, makes a call for the data, and the knowledge derived from that data, to become less distinct, and more easily navigated.
SageCite set about working closely with Sage Bionetworks to take some steps in investigating the development of disease network models, documenting the process, and adding support for citation.
Following a visit in November 2010 to the Sage Bioneworks base in Seattle, specific data sets and tools were provided by Brig Mecham at Sage Bionetworks to Peter Li. Using these data and tools, workflows were implemented in Taverna which captured and documented a particular stage of the disease model building process as part of the metaGEO project that is co-ordinated by Brig Mecham. These workflows, together with others being developed in the future, provided the basis for understanding the issues of data citation in network biology. These workflows are also an integral part of the demonstrator application that has been developed. The demonstrator shows how the DataCite service can be used for registering data that are generated from the building of disease network models. The workflows themselves are now shared through the MyExperiment environment.
The development and use of the demonstrator is described in these slides. A recording of the process will also be available shortly.
The registration of workflow data was implemented as a plugin for the Taverna workflow system. The plugin provided an activity which allows a data item to be associated with a DOI that is registered using the DataCite service. The decision on whether to register and cite workflow data is then made by the workflow builder during the development of the workflow. In addition, the DataCite can be used to associate the DOI with a web page which can be opened in a web browser to view the data item. For the purposes of software testing in the SageCite project, these web pages were created on a Google Sites web site.
Through the work on SageCite, a better understanding of how Sage Bionetwork’s predictive models of diseases are developed has been obtained. The complexity and work involved in building such models required close working between the two researchers at Sage Bionetworks and the SageCite project. As a result of the SageCite project, specific stages of the modelling process have been documented as Taverna workflows which are shared with the life sciences community using myExperiment. The SageCite project led to a collaboration on the metaGEO project with Brig Mecham from Sage Bionetworks who has developed tools for integrating gene expression data sets for meta-analyses of diseases, as described by Brig in his presentation at the Sage Congress. MetaGEO tools are now being used by Peter Li for studying blood cancers and diseases of inflammation at the University of Birmingham in their Systems Science for Health project.
This Monday, a number of thought leaders (and myself) will be gathering for two days of discussion on citation at the Data Citation Principles Workshop organised by IQSS at Harvard University.

I will be presenting some findings from our SageCite project, concentrating on examining the domain of disease modelling, and building on the thinking I started to develop at the JISC MRD meeting, for which I now have a graphic :
I am delighted to see that the line-up includes Gudmunder Thorisson presenting on ORCID and Max Wilkinson, our collaborator, talking about the DataCite International Initiative. Simon Hodson, the JISC programme manager for Research Data Management is also here for the meeting.
The list of discussion questions promises an interesting event.
The workshop is followed by an ORCID participant meeting on Wednesday: http://www.orcid.org/content/next-participant-meeting-may-18-boston
The presentation given at the Sage Congress about SageCite is now available as a pdf and as a stream. The rest of the presentations from the Congress are also available for viewing.
Starting from Slide 7 of Eric Schadt’s report on one of the breakout groups on sharing models, titled “Group B Incentives for sharing”, I want to highlight some observations on motivation for sharing and assigning credit. Two of the conclusions from the group were reported as “The number one outcome of our discussion was the desperate need to change how contributions are measured and get beyond journal articles being the only measure” and “Need to expand contributions to include: journals, DBs, datasets, curation, assertions, tools so that each are citable and scientists can get credit for these other types of activities” [slide 8]. Slide 9 provides a quick visual for how the measure of contributions needs to be re-balanced and some additional take-aways from the discussion are also summarised on the following slide. The stream for Eric’s report-back is also available, with the content on credit starting at about 04:28